Tingkatkan pengelasan ciri menggunakan projected quantum kernels
Anggaran penggunaan: 80 minit pada pemproses Heron r3 (NOTA: Ini hanya anggaran sahaja. Masa jalan sebenar anda mungkin berbeza.)
Dalam tutorial ini, kami menunjukkan cara menjalankan projected quantum kernel (PQK) dengan Qiskit pada dataset biologi dunia sebenar, berdasarkan kertas kerja Enhanced Prediction of CAR T-Cell Cytotoxicity with Quantum-Kernel Methods [1].
PQK ialah kaedah yang digunakan dalam pembelajaran mesin kuantum (QML) untuk mengekod data klasikal ke dalam ruang ciri kuantum dan mengunjurkannya semula ke domain klasikal, dengan menggunakan komputer kuantum untuk meningkatkan pemilihan ciri. Ia melibatkan pengekod data klasikal ke dalam keadaan kuantum menggunakan Circuit kuantum, biasanya melalui proses yang dipanggil pemetaan ciri, di mana data diubah ke dalam ruang Hilbert berdimensi tinggi. Aspek "projected" merujuk kepada pengekstrakan maklumat klasikal daripada keadaan kuantum, dengan mengukur observable tertentu, untuk membina matriks kernel yang boleh digunakan dalam algoritma berasaskan kernel klasikal seperti support vector machine. Pendekatan ini memanfaatkan kelebihan pengkomputeran sistem kuantum untuk berpotensi mencapai prestasi yang lebih baik pada tugas tertentu berbanding kaedah klasikal.
Tutorial ini juga mengandaikan kebiasaan umum dengan kaedah QML. Untuk penerokaan lanjut tentang QML, rujuk kursus Pembelajaran mesin kuantum dalam IBM Quantum Learning.
Keperluan
Sebelum memulakan tutorial ini, pastikan anda telah memasang perkara berikut:
- Qiskit SDK v2.0 atau lebih baru, dengan sokongan visualization
- Qiskit Runtime v0.40 atau lebih baru (
pip install qiskit-ibm-runtime) - Category encoders 2.8.1 (
pip install category-encoders) - NumPy 2.3.2 (
pip install numpy) - Pandas 2.3.2 (
pip install pandas) - Scikit-learn 1.7.1 (
pip install scikit-learn) - Tqdm 4.67.1 (
pip install tqdm)
Persediaan
# Added by doQumentation — required packages for this notebook
!pip install -q category-encoders numpy pandas qiskit qiskit-ibm-runtime scipy scikit-learn tqdm
import warnings
# Standard libraries
import os
import numpy as np
import pandas as pd
# Machine learning and data processing
import category_encoders as ce
from scipy.linalg import inv, sqrtm
from sklearn.metrics.pairwise import rbf_kernel
from sklearn.model_selection import GridSearchCV, StratifiedKFold
from sklearn.svm import SVC
# Qiskit and Qiskit Runtime
from qiskit import QuantumCircuit
from qiskit.circuit import ParameterVector
from qiskit.circuit.library import UnitaryGate, ZZFeatureMap
from qiskit.quantum_info import SparsePauliOp, random_unitary
from qiskit.transpiler import generate_preset_pass_manager
from qiskit_ibm_runtime import (
Batch,
EstimatorOptions,
EstimatorV2 as Estimator,
QiskitRuntimeService,
)
# Progress bar
import tqdm
warnings.filterwarnings("ignore")
Langkah 1: Petakan input klasikal kepada masalah kuantum
Penyediaan dataset
Dalam tutorial ini kami menggunakan dataset biologi dunia sebenar untuk tugasan pengelasan binari, yang dijana oleh Daniels et al. (2022) dan boleh dimuat turun dari bahan tambahan yang disertakan bersama kertas kerja tersebut. Data terdiri daripada sel T CAR, iaitu sel T yang direkayasa secara genetik dan digunakan dalam imunoterapi untuk merawat kanser tertentu. Sel T, sejenis sel imun, diubah suai di makmal untuk mengekspresikan reseptor antigen chimeric (CAR) yang menyasarkan protein tertentu pada sel kanser. Sel T yang diubah suai ini boleh mengenali dan memusnahkan sel kanser dengan lebih berkesan. Ciri-ciri data ialah motif sel T CAR, yang merujuk kepada komponen struktur atau fungsi khusus CAR yang direkayasa ke dalam sel T. Berdasarkan motif-motif ini, tugas kita adalah untuk meramal sitotoksisiti sel T CAR tertentu, melabelkannya sebagai toksik atau tidak toksik. Berikut menunjukkan fungsi pembantu untuk pra-proses dataset ini.
def preprocess_data(dir_root, args):
"""
Preprocess the training and test data.
"""
# Read from the csv files
train_data = pd.read_csv(
os.path.join(dir_root, args["file_train_data"]),
encoding="unicode_escape",
sep=",",
)
test_data = pd.read_csv(
os.path.join(dir_root, args["file_test_data"]),
encoding="unicode_escape",
sep=",",
)
# Fix the last motif ID
train_data[train_data == 17] = 14
train_data.columns = [
"Cell Number",
"motif",
"motif.1",
"motif.2",
"motif.3",
"motif.4",
"Nalm 6 Cytotoxicity",
]
test_data[test_data == 17] = 14
test_data.columns = [
"Cell Number",
"motif",
"motif.1",
"motif.2",
"motif.3",
"motif.4",
"Nalm 6 Cytotoxicity",
]
# Adjust motif at the third position
if args["filter_for_spacer_motif_third_position"]:
train_data = train_data[
(train_data["motif.2"] == 14) | (train_data["motif.2"] == 0)
]
test_data = test_data[
(test_data["motif.2"] == 14) | (test_data["motif.2"] == 0)
]
train_data = train_data[
args["motifs_to_use"] + [args["label_name"], "Cell Number"]
]
test_data = test_data[
args["motifs_to_use"] + [args["label_name"], "Cell Number"]
]
# Adjust motif at the last position
if not args["allow_spacer_motif_last_position"]:
last_motif = args["motifs_to_use"][len(args["motifs_to_use"]) - 1]
train_data = train_data[
(train_data[last_motif] != 14) & (train_data[last_motif] != 0)
]
test_data = test_data[
(test_data[last_motif] != 14) & (test_data[last_motif] != 0)
]
# Get the labels
train_labels = np.array(train_data[args["label_name"]])
test_labels = np.array(test_data[args["label_name"]])
# For the classification task use the threshold to binarize labels
train_labels[train_labels > args["label_binarization_threshold"]] = 1
train_labels[train_labels < 1] = args["min_label_value"]
test_labels[test_labels > args["label_binarization_threshold"]] = 1
test_labels[test_labels < 1] = args["min_label_value"]
# Reduce data to just the motifs of interest
train_data = train_data[args["motifs_to_use"]]
test_data = test_data[args["motifs_to_use"]]
# Get the class and motif counts
min_class = np.min(np.unique(np.concatenate([train_data, test_data])))
max_class = np.max(np.unique(np.concatenate([train_data, test_data])))
num_class = max_class - min_class + 1
num_motifs = len(args["motifs_to_use"])
print(str(max_class) + ":" + str(min_class) + ":" + str(num_class))
train_data = train_data - min_class
test_data = test_data - min_class
return (
train_data,
test_data,
train_labels,
test_labels,
num_class,
num_motifs,
)
def data_encoder(args, train_data, test_data, num_class, num_motifs):
"""
Use one-hot or binary encoding for classical data representation.
"""
if args["encoder"] == "one-hot":
# Transform to one-hot encoding
train_data = np.eye(num_class)[train_data]
test_data = np.eye(num_class)[test_data]
train_data = train_data.reshape(
train_data.shape[0], train_data.shape[1] * train_data.shape[2]
)
test_data = test_data.reshape(
test_data.shape[0], test_data.shape[1] * test_data.shape[2]
)
elif args["encoder"] == "binary":
# Transform to binary encoding
encoder = ce.BinaryEncoder()
base_array = np.unique(np.concatenate([train_data, test_data]))
base = pd.DataFrame(base_array).astype("category")
base.columns = ["motif"]
for motif_name in args["motifs_to_use"][1:]:
base[motif_name] = base.loc[:, "motif"]
encoder.fit(base)
train_data = encoder.transform(train_data.astype("category"))
test_data = encoder.transform(test_data.astype("category"))
train_data = np.reshape(
train_data.values, (train_data.shape[0], num_motifs, -1)
)
test_data = np.reshape(
test_data.values, (test_data.shape[0], num_motifs, -1)
)
train_data = train_data.reshape(
train_data.shape[0], train_data.shape[1] * train_data.shape[2]
)
test_data = test_data.reshape(
test_data.shape[0], test_data.shape[1] * test_data.shape[2]
)
else:
raise ValueError("Invalid encoding type.")
return train_data, test_data
Anda boleh menjalankan tutorial ini dengan menjalankan sel berikut, yang secara automatik mencipta struktur folder yang diperlukan dan memuat turun kedua-dua fail latihan dan ujian terus ke persekitaran anda. Jika anda sudah mempunyai fail-fail ini secara tempatan, langkah ini akan menimpanya dengan selamat untuk memastikan konsistensi versi.
## Download dataset
# Create data directory if it doesn't exist
!mkdir -p data_tutorial/pqk
# Download the training and test sets from the official Qiskit documentation repo
!wget -q --show-progress -O data_tutorial/pqk/train_data.csv \
https://raw.githubusercontent.com/Qiskit/documentation/main/datasets/tutorials/pqk/train_data.csv
!wget -q --show-progress -O data_tutorial/pqk/test_data.csv \
https://raw.githubusercontent.com/Qiskit/documentation/main/datasets/tutorials/pqk/test_data.csv
!wget -q --show-progress -O data_tutorial/pqk/projections_train.csv \
https://raw.githubusercontent.com/Qiskit/documentation/main/datasets/tutorials/pqk/projections_train.csv
!wget -q --show-progress -O data_tutorial/pqk/projections_test.csv \
https://raw.githubusercontent.com/Qiskit/documentation/main/datasets/tutorials/pqk/projections_test.csv
# Check the files have been downloaded
!echo "Dataset files downloaded:"
!ls -lh data_tutorial/pqk/*.csv
args = {
"file_train_data": "train_data.csv",
"file_test_data": "test_data.csv",
"motifs_to_use": ["motif", "motif.1", "motif.2", "motif.3"],
"label_name": "Nalm 6 Cytotoxicity",
"label_binarization_threshold": 0.62,
"filter_for_spacer_motif_third_position": False,
"allow_spacer_motif_last_position": True,
"min_label_value": -1,
"encoder": "one-hot",
}
dir_root = "./"
# Preprocess data
train_data, test_data, train_labels, test_labels, num_class, num_motifs = (
preprocess_data(dir_root=dir_root, args=args)
)
# Encode the data
train_data, test_data = data_encoder(
args, train_data, test_data, num_class, num_motifs
)
14:0:15
Kami juga mengubah dataset supaya diwakili sebagai untuk tujuan penskalaan.
# Change 1 to pi/2
angle = np.pi / 2
tmp = pd.DataFrame(train_data).astype("float64")
tmp[tmp == 1] = angle
train_data = tmp.values
tmp = pd.DataFrame(test_data).astype("float64")
tmp[tmp == 1] = angle
test_data = tmp.values
Kami mengesahkan saiz dan bentuk dataset latihan dan ujian.
print(train_data.shape, train_labels.shape)
print(test_data.shape, test_labels.shape)
(172, 60) (172,)
(74, 60) (74,)
Langkah 2: Optimumkan masalah untuk pelaksanaan perkakasan kuantum
Circuit kuantum
Kini kami membina peta ciri yang membenamkan dataset klasikal kami ke dalam ruang ciri berdimensi lebih tinggi. Untuk pembenaman ini, kami menggunakan ZZFeatureMap daripada Qiskit.
feature_dimension = train_data.shape[1]
reps = 24
insert_barriers = True
entanglement = "pairwise"
# ZZFeatureMap with linear entanglement and a repetition of 2
embed = ZZFeatureMap(
feature_dimension=feature_dimension,
reps=reps,
entanglement=entanglement,
insert_barriers=insert_barriers,
name="ZZFeatureMap",
)
embed.decompose().draw(output="mpl", style="iqp", fold=-1)

Pilihan pembenaman kuantum lain ialah ansatz evolusi Hamiltonian 1D-Heisenberg. Anda boleh melangkau bahagian ini jika anda ingin meneruskan dengan ZZFeatureMap.
feature_dimension = train_data.shape[1]
num_qubits = feature_dimension + 1
embed2 = QuantumCircuit(num_qubits)
num_trotter_steps = 6
pv_length = feature_dimension * num_trotter_steps
pv = ParameterVector("theta", pv_length)
# Add Haar random single qubit unitary to each qubit as initial state
np.random.seed(42)
seeds_unitary = np.random.randint(0, 100, num_qubits)
for i in range(num_qubits):
rand_gate = UnitaryGate(random_unitary(2, seed=seeds_unitary[i]))
embed2.append(rand_gate, [i])
def trotter_circ(feature_dimension, num_trotter_steps):
num_qubits = feature_dimension + 1
circ = QuantumCircuit(num_qubits)
# Even
for i in range(0, feature_dimension, 2):
circ.rzz(2 * pv[i] / num_trotter_steps, i, i + 1)
for i in range(0, feature_dimension, 2):
circ.rxx(2 * pv[i] / num_trotter_steps, i, i + 1)
for i in range(0, feature_dimension, 2):
circ.ryy(2 * pv[i] / num_trotter_steps, i, i + 1)
# Odd
for i in range(1, feature_dimension, 2):
circ.rzz(2 * pv[i] / num_trotter_steps, i, i + 1)
for i in range(1, feature_dimension, 2):
circ.rxx(2 * pv[i] / num_trotter_steps, i, i + 1)
for i in range(1, feature_dimension, 2):
circ.ryy(2 * pv[i] / num_trotter_steps, i, i + 1)
return circ
# Hamiltonian evolution ansatz
for step in range(num_trotter_steps):
circ = trotter_circ(feature_dimension, num_trotter_steps)
if step % 2 == 0:
embed2 = embed2.compose(circ)
else:
reverse_circ = circ.reverse_ops()
embed2 = embed2.compose(reverse_circ)
embed2.draw(output="mpl", style="iqp", fold=-1)
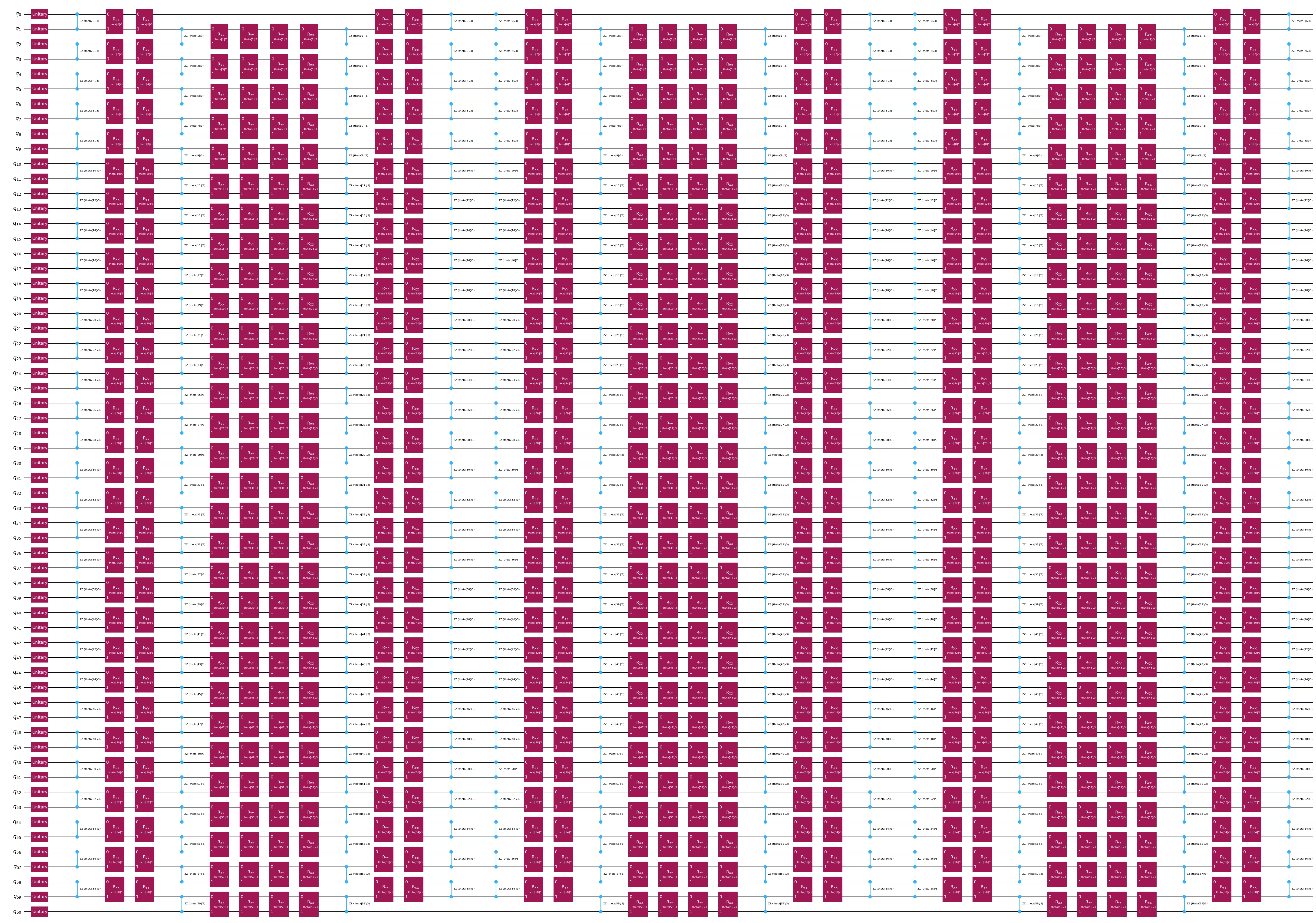
Langkah 3: Laksanakan menggunakan Qiskit primitives
Ukur 1-RDM
Blok pembina utama projected quantum kernels ialah matriks ketumpatan tereduksi (RDM), yang diperoleh melalui pengukuran projektif peta ciri kuantum. Dalam langkah ini, kami memperoleh semua matriks ketumpatan tereduksi satu-Qubit (1-RDM), yang kemudiannya akan dimasukkan ke dalam fungsi kernel eksponen klasikal.
Mari kita lihat cara mengira 1-RDM diberi satu titik data dari dataset sebelum kami menjalani semua data. 1-RDM ialah koleksi pengukuran satu-Qubit operator Pauli X, Y dan Z pada semua Qubit. Ini kerana satu-Qubit RDM boleh dinyatakan sepenuhnya sebagai:
Pertama kami memilih Backend yang hendak digunakan.
service = QiskitRuntimeService()
backend = service.least_busy(
operational=True, simulator=False, min_num_qubits=133
)
target = backend.target
Kemudian kami menjalankan Circuit kuantum dan mengukur unjuran. Perhatikan bahawa kami menghidupkan mitigasi ralat, termasuk Zero Noise Extrapolation (ZNE).
# Let's select the ZZFeatureMap embedding for this example
qc = embed
num_qubits = feature_dimension
# Identity operator on all qubits
id = "I" * num_qubits
# Let's select the first training datapoint as an example
parameters = train_data[0]
# Bind parameter to the circuit and simplify it
qc_bound = qc.assign_parameters(parameters)
transpiler = generate_preset_pass_manager(
optimization_level=3, basis_gates=["u3", "cz"]
)
transpiled_circuit = transpiler.run(qc_bound)
# Transpile for hardware
transpiler = generate_preset_pass_manager(optimization_level=3, target=target)
transpiled_circuit = transpiler.run(transpiled_circuit)
# We group all commuting observables
# These groups are the Pauli X, Y and Z operators on individual qubits
observables_x = [
SparsePauliOp(id[:i] + "X" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_y = [
SparsePauliOp(id[:i] + "Y" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_z = [
SparsePauliOp(id[:i] + "Z" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
# We define the primitive unified blocs (PUBs) consisting of the embedding circuit,
# set of observables and the circuit parameters
pub_x = (transpiled_circuit, observables_x)
pub_y = (transpiled_circuit, observables_y)
pub_z = (transpiled_circuit, observables_z)
# Experiment options for error mitigation
num_randomizations = 300
shots_per_randomization = 100
noise_factors = [1, 3, 5]
experimental_opts = {}
experimental_opts["resilience"] = {
"measure_mitigation": True,
"zne_mitigation": True,
"zne": {
"noise_factors": noise_factors,
"amplifier": "gate_folding",
"extrapolated_noise_factors": [0] + noise_factors,
},
}
experimental_opts["twirling"] = {
"num_randomizations": num_randomizations,
"shots_per_randomization": shots_per_randomization,
"strategy": "active-accum",
}
# We define and run the estimator to obtain <X>, <Y> and <Z> on all qubits
estimator = Estimator(mode=backend, options=experimental_opts)
job = estimator.run([pub_x, pub_y, pub_z])
Seterusnya kami mendapatkan semula keputusan.
job_result_x = job.result()[0].data.evs
job_result_y = job.result()[1].data.evs
job_result_z = job.result()[2].data.evs
print(job_result_x)
print(job_result_y)
print(job_result_z)
[ 3.67865951e-03 1.01158571e-02 -3.95790878e-02 6.33984326e-03
1.86035759e-02 -2.91533268e-02 -1.06374793e-01 4.48873518e-18
4.70201764e-02 3.53997968e-02 2.53130819e-02 3.23903401e-02
6.06327843e-03 1.16313667e-02 -1.12387504e-02 -3.18457725e-02
-4.16445718e-04 -1.45609602e-03 -4.21737114e-01 2.83705669e-02
6.91332890e-03 -7.45363001e-02 -1.20139326e-02 -8.85566135e-02
-3.22648394e-02 -3.24228074e-02 6.20431299e-04 3.04225434e-03
5.72795792e-03 1.11288428e-02 1.50395861e-01 9.18380197e-02
1.02553163e-01 2.98312847e-02 -3.30298912e-01 -1.13979648e-01
4.49159340e-03 8.63861493e-02 3.05666566e-02 2.21463145e-04
1.45946735e-02 8.54537275e-03 -8.09805979e-02 -2.92608104e-02
-3.91243644e-02 -3.96632760e-02 -1.41187613e-01 -1.07363243e-01
1.81089440e-02 2.70778895e-02 1.45139414e-02 2.99480458e-02
4.99137134e-02 7.08789852e-02 4.30565759e-02 8.71287156e-02
1.04334798e-01 7.72191962e-02 7.10059720e-02 1.04650403e-01]
[-7.31765102e-05 7.42669174e-03 9.82277344e-03 5.92638249e-02
4.24120486e-02 -9.06473416e-03 4.55057675e-03 8.43494094e-03
6.92097339e-02 -6.82234424e-02 6.13509008e-02 3.94200491e-02
-1.24037979e-02 1.01976642e-01 7.90538600e-03 -7.19726160e-02
-1.19501703e-16 -1.03796614e-02 7.37382463e-02 1.97238568e-01
-3.59250635e-02 -2.67554009e-02 3.55010633e-02 7.68877990e-02
6.50677589e-05 -6.59298767e-03 -1.23719487e-02 -6.41938151e-02
1.95603072e-02 -2.48448551e-02 5.17784810e-02 -5.93767100e-02
3.11897681e-02 -3.91959720e-18 -4.47769148e-03 1.39202197e-01
-6.56387523e-02 -5.85665483e-02 9.52905894e-03 -8.61460731e-02
3.91790656e-02 -1.27544375e-01 1.63712244e-01 3.36816934e-04
2.26230028e-02 -2.45023393e-05 4.95635588e-03 1.44779564e-01
3.71625177e-02 3.65675948e-03 2.83694017e-02 -7.10500602e-02
-1.15467702e-01 6.21712129e-03 -4.80958959e-02 2.21021066e-02
7.99062499e-02 -1.87164076e-02 -3.67100369e-02 -2.38923731e-02]
[ 6.85871605e-01 5.07725024e-01 8.71024642e-03 3.34823455e-02
4.58684961e-02 9.44384189e-17 -4.46829296e-02 -2.91296778e-02
4.15466461e-02 2.89628330e-02 1.88624017e-03 5.37110446e-02
2.59579053e-03 1.39327071e-02 -2.90781778e-02 5.07209866e-03
5.83403000e-02 2.60764440e-02 4.45999706e-17 -6.66701417e-03
3.03215873e-01 2.26172533e-02 2.43105960e-02 4.98861041e-18
-2.45530791e-02 6.26940708e-02 1.21058073e-02 2.76675948e-04
2.63980996e-02 2.58302364e-02 7.47856723e-02 8.42728943e-02
5.70989097e-02 6.92955086e-02 -5.68313712e-03 1.32199452e-01
8.90511238e-02 -3.45204621e-02 -1.05445836e-01 6.03864150e-03
2.16291384e-02 8.22303162e-03 1.00856715e-02 6.28973151e-02
6.26727169e-02 6.15399206e-02 9.67320897e-02 1.03045269e-16
1.79688783e-01 -1.59960520e-02 -1.15422952e-02 9.60200470e-03
6.58396672e-02 7.78329830e-03 6.53226955e-02 2.45778685e-03
4.36694753e-03 5.75098762e-03 -2.48896201e-02 8.33740755e-05]
Kami mencetak saiz Circuit dan kedalaman Gate dua-Qubit.
print(f"qubits: {qc.num_qubits}")
print(
f"2q-depth: {transpiled_circuit.depth(lambda x: x.operation.num_qubits==2)}"
)
print(
f"2q-size: {transpiled_circuit.size(lambda x: x.operation.num_qubits==2)}"
)
print(f"Operator counts: {transpiled_circuit.count_ops()}")
transpiled_circuit.draw("mpl", fold=-1, style="clifford", idle_wires=False)
qubits: 60
2q-depth: 64
2q-size: 1888
Operator counts: OrderedDict({'rz': 6016, 'sx': 4576, 'cz': 1888, 'x': 896, 'barrier': 31})

Kini kami boleh membuat gelung ke atas keseluruhan dataset latihan untuk memperoleh semua 1-RDM.
Kami juga menyediakan keputusan daripada eksperimen yang kami jalankan pada perkakasan kuantum. Anda boleh sama ada menjalankan latihan sendiri dengan menetapkan bendera di bawah kepada True, atau menggunakan keputusan unjuran yang kami sediakan.
# Set this to True if you want to run the training on hardware
run_experiment = False
# Identity operator on all qubits
id = "I" * num_qubits
# projections_train[i][j][k] will be the expectation value of the j-th Pauli operator (0: X, 1: Y, 2: Z)
# of datapoint i on qubit k
projections_train = []
jobs_train = []
# Experiment options for error mitigation
num_randomizations = 300
shots_per_randomization = 100
noise_factors = [1, 3, 5]
experimental_opts = {}
experimental_opts["resilience"] = {
"measure_mitigation": True,
"zne_mitigation": True,
"zne": {
"noise_factors": noise_factors,
"amplifier": "gate_folding",
"return_all_extrapolated": True,
"return_unextrapolated": True,
"extrapolated_noise_factors": [0] + noise_factors,
},
}
experimental_opts["twirling"] = {
"num_randomizations": num_randomizations,
"shots_per_randomization": shots_per_randomization,
"strategy": "active-accum",
}
options = EstimatorOptions(experimental=experimental_opts)
if run_experiment:
with Batch(backend=backend):
for i in tqdm.tqdm(
range(len(train_data)), desc="Training data progress"
):
# Get training sample
parameters = train_data[i]
# Bind parameter to the circuit and simplify it
qc_bound = qc.assign_parameters(parameters)
transpiler = generate_preset_pass_manager(
optimization_level=3, basis_gates=["u3", "cz"]
)
transpiled_circuit = transpiler.run(qc_bound)
# Transpile for hardware
transpiler = generate_preset_pass_manager(
optimization_level=3, target=target
)
transpiled_circuit = transpiler.run(transpiled_circuit)
# We group all commuting observables
# These groups are the Pauli X, Y and Z operators on individual qubits
observables_x = [
SparsePauliOp(id[:i] + "X" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_y = [
SparsePauliOp(id[:i] + "Y" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_z = [
SparsePauliOp(id[:i] + "Z" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
# We define the primitive unified blocs (PUBs) consisting of the embedding circuit,
# set of observables and the circuit parameters
pub_x = (transpiled_circuit, observables_x)
pub_y = (transpiled_circuit, observables_y)
pub_z = (transpiled_circuit, observables_z)
# We define and run the estimator to obtain <X>, <Y> and <Z> on all qubits
estimator = Estimator(options=options)
job = estimator.run([pub_x, pub_y, pub_z])
jobs_train.append(job)
Training data progress: 100%|██████████| 172/172 [13:03<00:00, 4.55s/it]
Setelah kerja selesai, kami boleh mendapatkan semula keputusan.
if run_experiment:
for i in tqdm.tqdm(
range(len(train_data)), desc="Retrieving training data results"
):
# Completed job
job = jobs_train[i]
# Job results
job_result_x = job.result()[0].data.evs
job_result_y = job.result()[1].data.evs
job_result_z = job.result()[2].data.evs
# Record <X>, <Y> and <Z> on all qubits for the current datapoint
projections_train.append([job_result_x, job_result_y, job_result_z])
Kami mengulangi ini untuk set ujian.
# Identity operator on all qubits
id = "I" * num_qubits
# projections_test[i][j][k] will be the expectation value of the j-th Pauli operator (0: X, 1: Y, 2: Z)
# of datapoint i on qubit k
projections_test = []
jobs_test = []
# Experiment options for error mitigation
num_randomizations = 300
shots_per_randomization = 100
noise_factors = [1, 3, 5]
experimental_opts = {}
experimental_opts["resilience"] = {
"measure_mitigation": True,
"zne_mitigation": True,
"zne": {
"noise_factors": noise_factors,
"amplifier": "gate_folding",
"return_all_extrapolated": True,
"return_unextrapolated": True,
"extrapolated_noise_factors": [0] + noise_factors,
},
}
experimental_opts["twirling"] = {
"num_randomizations": num_randomizations,
"shots_per_randomization": shots_per_randomization,
"strategy": "active-accum",
}
options = EstimatorOptions(experimental=experimental_opts)
if run_experiment:
with Batch(backend=backend):
for i in tqdm.tqdm(range(len(test_data)), desc="Test data progress"):
# Get test sample
parameters = test_data[i]
# Bind parameter to the circuit and simplify it
qc_bound = qc.assign_parameters(parameters)
transpiler = generate_preset_pass_manager(
optimization_level=3, basis_gates=["u3", "cz"]
)
transpiled_circuit = transpiler.run(qc_bound)
# Transpile for hardware
transpiler = generate_preset_pass_manager(
optimization_level=3, target=target
)
transpiled_circuit = transpiler.run(transpiled_circuit)
# We group all commuting observables
# These groups are the Pauli X, Y and Z operators on individual qubits
observables_x = [
SparsePauliOp(id[:i] + "X" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_y = [
SparsePauliOp(id[:i] + "Y" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
observables_z = [
SparsePauliOp(id[:i] + "Z" + id[(i + 1) :]).apply_layout(
transpiled_circuit.layout
)
for i in range(num_qubits)
]
# We define the primitive unified blocs (PUBs) consisting of the embedding circuit,
# set of observables and the circuit parameters
pub_x = (transpiled_circuit, observables_x)
pub_y = (transpiled_circuit, observables_y)
pub_z = (transpiled_circuit, observables_z)
# We define and run the estimator to obtain <X>, <Y> and <Z> on all qubits
estimator = Estimator(options=options)
job = estimator.run([pub_x, pub_y, pub_z])
jobs_test.append(job)
Test data progress: 100%|██████████| 74/74 [00:13<00:00, 5.56it/s]
Kami boleh mendapatkan semula keputusan seperti sebelumnya.
if run_experiment:
for i in tqdm.tqdm(
range(len(test_data)), desc="Retrieving test data results"
):
# Completed job
job = jobs_test[i]
# Job results
job_result_x = job.result()[0].data.evs
job_result_y = job.result()[1].data.evs
job_result_z = job.result()[2].data.evs
# Record <X>, <Y> and <Z> on all qubits for the current datapoint
projections_test.append([job_result_x, job_result_y, job_result_z])
Langkah 4: Pasca-proses dan kembalikan keputusan dalam format klasikal yang dikehendaki
Takrifkan projected quantum kernel
Projected quantum kernel ditakrifkan dengan fungsi kernel berikut:
Dalam persamaan di atas, ialah hiperparameter yang boleh dilaras. ialah entri matriks kernel .
Menggunakan takrif 1-RDM, kita dapat melihat bahawa sebutan individu dalam fungsi kernel boleh dinilai sebagai , di mana . Nilai jangkaan ini adalah tepat seperti yang kami ukur di atas.
Dengan menggunakan scikit-learn, kita sebenarnya boleh mengira kernel dengan lebih mudah. Ini disebabkan oleh fungsi kernel radial basis ('rbf') yang tersedia: . Pertama, kita hanya perlu membentuk semula dataset latihan dan ujian yang diunjurkan baharu ke dalam tatasusunan dua dimensi.
Perhatikan bahawa menjalani keseluruhan dataset boleh mengambil masa sekitar 80 minit pada QPU. Untuk memastikan tutorial selebihnya mudah dilaksanakan, kami turut menyediakan unjuran daripada eksperimen yang telah dijalankan sebelumnya (yang disertakan dalam fail yang anda muat turun dalam blok kod Download dataset). Jika anda telah menjalankan latihan sendiri, anda boleh meneruskan tutorial dengan keputusan anda sendiri.
if run_experiment:
projections_train = np.array(projections_train).reshape(
len(projections_train), -1
)
projections_test = np.array(projections_test).reshape(
len(projections_test), -1
)
else:
projections_train = np.loadtxt("projections_train.txt")
projections_test = np.loadtxt("projections_test.txt")
Support Vector Machine (SVM)
Kini kami boleh menjalankan SVM klasikal pada kernel yang telah dikira ini, dan menggunakan kernel antara set ujian dan latihan untuk membuat ramalan.
# Range of 'C' and 'gamma' values as SVC hyperparameters
C_range = [0.001, 0.005, 0.007]
C_range.extend([x * 0.01 for x in range(1, 11)])
C_range.extend([x * 0.25 for x in range(1, 60)])
C_range.extend(
[
20,
50,
100,
200,
500,
700,
1000,
1100,
1200,
1300,
1400,
1500,
1700,
2000,
]
)
gamma_range = ["auto", "scale", 0.001, 0.005, 0.007]
gamma_range.extend([x * 0.01 for x in range(1, 11)])
gamma_range.extend([x * 0.25 for x in range(1, 60)])
gamma_range.extend([20, 50, 100])
param_grid = dict(C=C_range, gamma=gamma_range)
# Support vector classifier
svc = SVC(kernel="rbf")
# Define the cross validation
cv = StratifiedKFold(n_splits=10)
# Grid search for hyperparameter tuning (q: quantum)
grid_search_q = GridSearchCV(
svc, param_grid, cv=cv, verbose=1, n_jobs=-1, scoring="f1_weighted"
)
grid_search_q.fit(projections_train, train_labels)
# Best model with best parameters
best_svc_q = grid_search_q.best_estimator_
print(
f"The best parameters are {grid_search_q.best_params_} with a score of {grid_search_q.best_score_:.4f}"
)
# Test accuracy
accuracy_q = best_svc_q.score(projections_test, test_labels)
print(f"Test accuracy with best model: {accuracy_q:.4f}")
Fitting 10 folds for each of 6622 candidates, totalling 66220 fits
The best parameters are {'C': 8.5, 'gamma': 0.01} with a score of 0.6980
Test accuracy with best model: 0.8108
Penanda aras klasikal
Kami boleh menjalankan SVM klasikal dengan fungsi radial basis sebagai kernel tanpa melakukan unjuran kuantum. Keputusan ini ialah penanda aras klasikal kami.
# Support vector classifier
svc = SVC(kernel="rbf")
# Grid search for hyperparameter tuning (c: classical)
grid_search_c = GridSearchCV(
svc, param_grid, cv=cv, verbose=1, n_jobs=-1, scoring="f1_weighted"
)
grid_search_c.fit(train_data, train_labels)
# Best model with best parameters
best_svc_c = grid_search_c.best_estimator_
print(
f"The best parameters are {grid_search_c.best_params_} with a score of {grid_search_c.best_score_:.4f}"
)
# Test accuracy
accuracy_c = best_svc_c.score(test_data, test_labels)
print(f"Test accuracy with best model: {accuracy_c:.4f}")
Fitting 10 folds for each of 6622 candidates, totalling 66220 fits
The best parameters are {'C': 10.75, 'gamma': 0.04} with a score of 0.7830
Test accuracy with best model: 0.7432
Lampiran: Sahkan potensi kelebihan kuantum dataset dalam tugasan pembelajaran
Tidak semua dataset menawarkan potensi kelebihan daripada penggunaan PQK. Terdapat beberapa had teori yang boleh digunakan sebagai ujian awal untuk melihat sama ada dataset tertentu boleh mendapat manfaat daripada PQK. Untuk mengukur ini, pengarang Power of data in quantum machine learning [2] mentakrifkan kuantiti yang dirujuk sebagai kerumitan model klasikal dan kuantum serta pemisahan geometri model klasikal dan kuantum. Untuk mengharapkan potensi kelebihan kuantum daripada PQK, pemisahan geometri antara kernel klasikal dan kernel yang diunjurkan secara kuantum hendaklah lebih kurang pada orde , di mana ialah bilangan sampel latihan. Jika syarat ini dipenuhi, kita teruskan dengan memeriksa kerumitan model. Jika kerumitan model klasikal adalah pada orde manakala kerumitan model yang diunjurkan secara kuantum jauh lebih kecil daripada , kita boleh mengharapkan potensi kelebihan daripada PQK. Pemisahan geometri ditakrifkan seperti berikut (F19 dalam [2]):
# Gamma values used in best models above
gamma_c = grid_search_c.best_params_["gamma"]
gamma_q = grid_search_q.best_params_["gamma"]
# Regularization parameter used in the best classical model above
C_c = grid_search_c.best_params_["C"]
l_c = 1 / C_c
# Classical and quantum kernels used above
K_c = rbf_kernel(train_data, train_data, gamma=gamma_c)
K_q = rbf_kernel(projections_train, projections_train, gamma=gamma_q)
# Intermediate matrices in the equation
K_c_sqrt = sqrtm(K_c)
K_q_sqrt = sqrtm(K_q)
K_c_inv = inv(K_c + l_c * np.eye(K_c.shape[0]))
K_multiplication = (
K_q_sqrt @ K_c_sqrt @ K_c_inv @ K_c_inv @ K_c_sqrt @ K_q_sqrt
)
# Geometric separation
norm = np.linalg.norm(K_multiplication, ord=np.inf)
g_cq = np.sqrt(norm)
print(
f"Geometric separation between classical and quantum kernels is {g_cq:.4f}"
)
print(np.sqrt(len(train_data)))
Geometric separation between classical and quantum kernels is 1.5440
13.114877048604
Kerumitan model ditakrifkan seperti berikut (M1 dalam [2]):
# Model complexity of the classical kernel
# Number of training data
N = len(train_data)
# Predicted labels
pred_labels = best_svc_c.predict(train_data)
pred_matrix = np.outer(pred_labels, pred_labels)
# Intermediate terms
K_c_inv = inv(K_c + l_c * np.eye(K_c.shape[0]))
# First term
first_sum = np.sum((K_c_inv @ K_c_inv) * pred_matrix)
first_term = l_c * np.sqrt(first_sum / N)
# Second term
second_sum = np.sum((K_c_inv @ K_c @ K_c_inv) * pred_matrix)
second_term = np.sqrt(second_sum / N)
# Model complexity
s_c = first_term + second_term
print(f"Classical model complexity is {s_c:.4f}")
Classical model complexity is 1.3578
# Model complexity of the projected quantum kernel
# Number of training data
N = len(projections_train)
# Predicted labels
pred_labels = best_svc_q.predict(projections_train)
pred_matrix = np.outer(pred_labels, pred_labels)
# Regularization parameter used in the best classical model above
C_q = grid_search_q.best_params_["C"]
l_q = 1 / C_q
# Intermediate terms
K_q_inv = inv(K_q + l_q * np.eye(K_q.shape[0]))
# First term
first_sum = np.sum((K_q_inv @ K_q_inv) * pred_matrix)
first_term = l_q * np.sqrt(first_sum / N)
# Second term
second_sum = np.sum((K_q_inv @ K_q @ K_q_inv) * pred_matrix)
second_term = np.sqrt(second_sum / N)
# Model complexity
s_q = first_term + second_term
print(f"Quantum model complexity is {s_q:.4f}")
Quantum model complexity is 1.5806
Rujukan
- Utro, Filippo, et al. "Enhanced Prediction of CAR T-Cell Cytotoxicity with Quantum-Kernel Methods." arXiv preprint arXiv:2507.22710 (2025).
- Huang, Hsin-Yuan, et al. "Power of data in quantum machine learning." Nature communications 12.1 (2021): 2631.
- Daniels, Kyle G., et al. "Decoding CAR T cell phenotype using combinatorial signaling motif libraries and machine learning." Science 378.6625 (2022): 1194-1200.